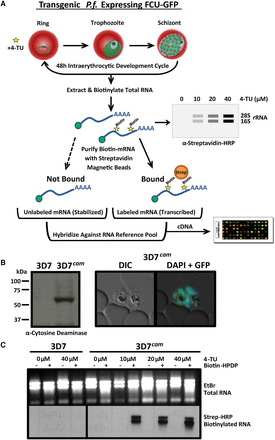
Engineering P. falciparum to salvage pyrimidines and generate thiol-modified RNAs. (A) Schematic of 4-TU biosynthetic mRNA capture method. Transgenic P. falciparum expressing a fusion gene containing cytosine deaminase/uracil phosphoribosyltransferase tagged with GFP (FCU-GFP) under the control of the calmodulin (cam) promoter (3D7cam) enables 4-TU salvage and incorporation into RNA. Thiolated-RNA can be biotinylated and detected by Northern blot or affinity purified by streptavidin magnetic beads for analysis by DNA microarray (or RNA-seq). (B) Expression of FCU-GFP in 3D7cam parasites was verified by Western blot when probed with anti-yeast cytosine deaminase and by live fluorescence microscopy: (green) GFP; (blue) nuclear DNA stained with DAPI. (C) Both wild-type and 3D7cam parasites were grown for 12 h in the presence of increasing 4-TU concentrations. The specificity of RNA thiol-incorporation and biotinylation was assessed by running 2 µg of each RNA sample with and without EZ-link Biotin-HPDP incubation (top). Total RNA was transferred to a nylon membrane and probed with streptavidin-HRP to detect biotinylated RNAs (bottom).











