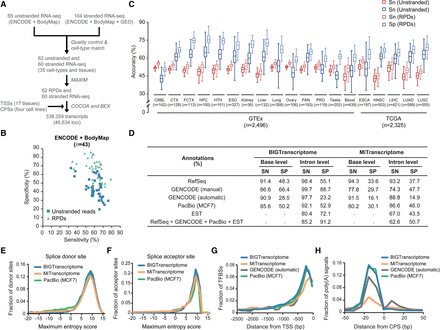
Comprehensive human transcriptome map. (A) A schematic flow for the reconstruction of the BIGTranscriptome map using large-scale RNA-seq samples from human cell lines, ENCODE, and Human BodyMap 2.0 Projects. (B) Accuracies of unstranded (blue) and RPD assemblies (mint) from the ENCODE and Human BodyMap 2.0 projects. (C) Sensitivities (red) and specificities (blue) of unstranded assemblies (solid line box) and RPD assemblies (dotted line box) are shown in box plots. The unstranded RNA-seq data are from GTEx (14 tissues) and TCGA Project (five tumor types). The numbers (n) indicate the sample numbers in each group. (CRBL) Brain cerebellum, (CTX) brain cortex, (FCTX) brain frontal cortex, (HPC) brain hippocampus, (HTH) brain hypothalamus, (ESO) esophagus-mucosa, (PAN) pancreas, (PRO) prostate, (ESCA) esophageal carcinoma, (HNSC) head and neck squamous cell carcinoma, (LIHC) liver hepatocellular carcinoma, (LUAD) lung adenocarcinoma, and (LUSC) lung squamous cell carcinoma. (D) Shown are the accuracies of BIGTranscriptome and MiTranscriptome at the base and intron levels based on four different sets of annotations (RefSeq, manual and automatic GENCODE, PacBio, and EST), and a combined set of annotations. (SN) Sensitivity, (SP) specificity. (E,F) Maximum entropy scores of the putative splice donor sites (E) and of putative splice acceptor sites (F). Blue lines are from BIGTranscriptome, green lines are from PacBio assembly, and orange lines are from MiTranscriptome. (G) The fraction of TFBSs upstream of the 5′ end of BIGTranscriptome transcripts (blue) was compared to those of MiTranscriptome (orange), GENCODE (automatic) (black), and PacBio assembly (green). (H) The fraction of the closest poly(A) signals, AAUAAA, in the region just upstream of the 3′ end of BIGTranscriptome annotations (blue) compared to those of MiTranscriptome (orange), GENCODE (automatic) (black), and PacBio assembly (green).











