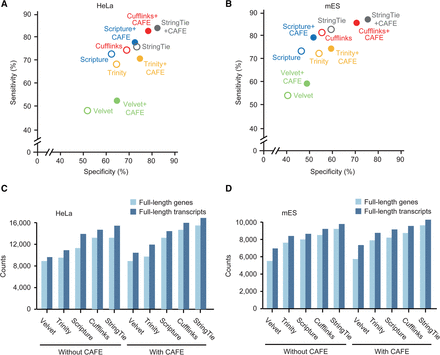
Benchmarking other base assemblers. (A,B) The accuracies of combined transcriptome assemblies (solid circles) reconstructed by CAFE with base assemblers and of the original transcriptome assemblies (open circles) reconstructed by respective base assemblers, such as Cufflinks (red), Scripture (blue), StringTie (gray), Velvet (green), and Trinity (yellow), in HeLa (A) and mES cells (B). The accuracies of the original assemblies were calculated by averaging the accuracies of stranded and unstranded assemblies reconstructed by each base assembler. Velvet and Trinity were used as de novo assemblers, and Scripture, StringTie, and Cufflinks were used as reference-based assemblers. (C,D) The numbers of full-length genes (light blue) and transcripts (blue) in the coassemblies were compared to those in the original assemblies from HeLa (C) and mES cells (D). For the original assemblies, the higher number of full-length genes in the stranded and unstranded original assemblies was chosen.











