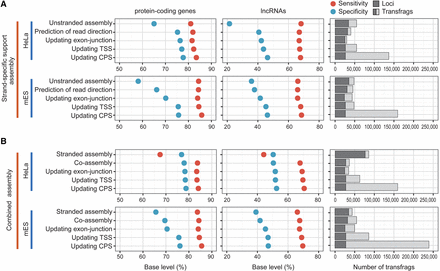
Step-wise evaluation of transcriptomes reassembled by CAFE. (A) Shown are the accuracies and sizes of strand-specific support transcriptomes (RPD assembly) at each step of CAFE in HeLa (top) and mES cells (bottom). The sensitivity (red solid circle) and specificity (blue) of the assemblies are measured by comparing to GENCODE protein-coding genes (left panel) and lncRNAs (middle panel). The number of assembled transfrags and their loci are indicated at each step (right panel). (B) Shown are the accuracies and sizes of combined transcriptome assemblies of both stranded reads and RPDs. The low sensitivity of the stranded assembly from HeLa cells is presumably because the stranded reads are of the single-end type and are 36 or 72 nt long. Otherwise, as in A.











