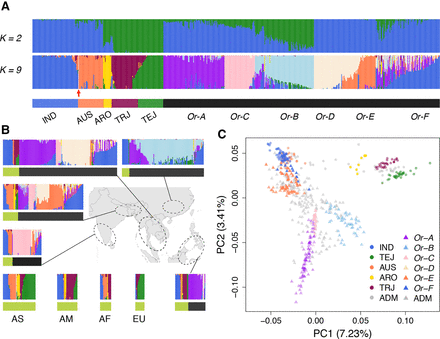
Population structure of the rice primary gene pool, including O. sativa and O. rufipogon. (A) Clustering using NGSadmix assuming K = 2 and K = 9. At K = 2, the samples are divided into indica and japonica components. At K = 9, five subgroups of domesticated rice are recovered, and four unique components of wild rice are identified. The color bars beneath the clusters denote the subgroup assignments. The red arrow points to the misidentified domesticated accession (GSOR311586), which was confirmed to have wild rice ancestry. The abbreviations of subgroups in cultivated rice are as follows: (ADM) admixture; (IND) indica; (AUS) aus; (ARO) aromatic; (TRJ) tropical japonica; (TEJ) temperate japonica. (B) Geographic distribution of rice samples. South and Southeast Asia, which are the major habitats for wild rice and also major rice cultivation areas, are shown on the map. The area was divided into five regions: (1) South Asia; (2) Ganges Basin; (3) Indochina Peninsula; (4) China; (5) Archipelago countries. The color code of the bar beneath the clustering plot indicates cultivated (green) and wild (black) rice. The abbreviations at the bottom left are as follows: (AS) Asia; all rice samples from Asia but not shown on the map are included in this category; (AM) America; (AF) Africa; (EU) Europe. (C) PCA of the combined population with wild (triangle) and cultivated (dot) rice samples. The abbreviation codes are the same as those in A.











