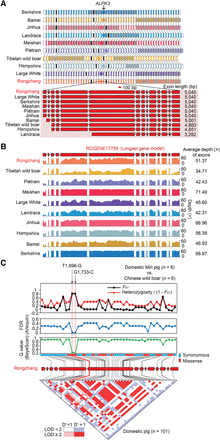
Details of assembled ALPK3 gene and selected variants. (A) Structure of assembled ALPK3. (Top panel) The interassembly collinear genes (colored rectangles) among 10 assemblies are linked by gray lines, and the genes not present in all 10 assemblies are marked in black. ALPK3 is denoted by a circle. Different scaffolds are shown as alternating white and gray backgrounds. (Bottom panel) Comparison of structure of ALPK3 among the 10 assemblies. Boxes and lines indicate exons and introns, respectively. (B) Coverage and depth for the longest gene model of ALPK3 (Gene ID: RCGENE17759) by cross-mapping reads from paired-end DNA libraries (insert sizes of 180 and 500 bp) of the 10 assemblies. The higher coverage depth (≥30×) suggests slightly different structures of ALPK3, which is attributable to limitations of short read assembly; as such, the longest gene model is considered more reliable and used for subsequent analyses. (C) Two selected missense mutations (T1,696-G and G1,733-C) in ALPK3 between Chinese wild boars (n = 6) and domestic Min pigs (n = 6). (Top panels) FST and heterozygosity/(1−FST), FDR (Arlequin), and Q-values (BayeScan) are plotted for 45 coding SNPs (18 missenses and 27 synonymous mutations). (Bottom panels) LD pattern of 45 SNPs in 101 domestic pigs from China (n = 41), North America (n = 12), and Europe (n = 48). Squares shaded in pink or red indicate significant LD between SNP pairs (bright red indicates pairwise D′ = 1), white squares indicate no evidence of significant LD, and blue squares indicate pairwise D′ = 1 without statistical significance. The adjacent T1,696-G and G1,733-C are closely linked (D′ = 1, r2 = 0.975, LOD = 41.6).











