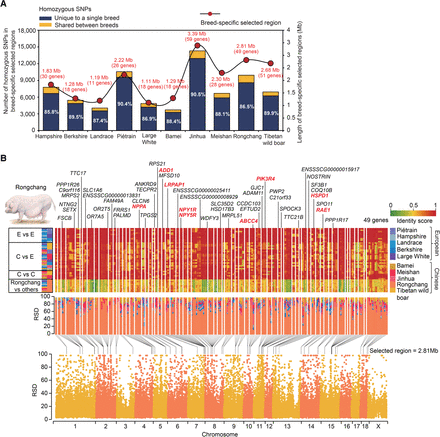
Identification of breed-specific selective sweeps. (A) Number of homozygous SNPs in breed-specific selected regions. Of 74.21 k homozygous SNPs in 20.10 Mb selected regions, 65.75 k (88.60%) were unique to a particular breed, which was highly concentrated in a small fraction (0.79%) of the genome and likely contributed to diversifying selection. (B) Selective sweep regions identified in the Rongchang pig. (Top panels, top half) Genes residing within or in the vicinity (±5 kb) of the selected regions are presented for each chromosome and ordered according to their locations. (Top panels, lower half) Degree of haplotype sharing of selected regions in pairwise comparisons among the 10 breeds. Homozygous SNP frequencies in individual breeds were used to calculate identity scores in 10-kb windows. Boxes (left) indicate pairwise comparison presented on that row (E, European pigs; C, Chinese pigs) according to the color assigned to each pig breed (right). Heat map colors indicate identity scores. (Middle panel) Percentage stacked column showing RSD values in the Rongchang-specific selected regions across 10 breeds sequenced. Rongchang showed predominantly higher RSD values than other breeds, indicating that only this breed has SNPs compared to the reference genome in this region. (Bottom half) RSD in 10-kb windows for Rongchang plotted along chromosomes. Black lines indicate selected regions (FDR < 0.05). Nine selected genes orthologous to the mammalian fat deposition genes are marked in red.











