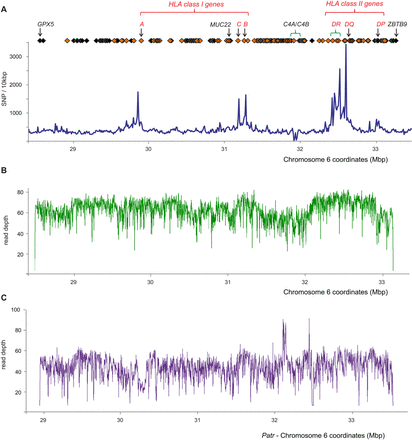
The target MHC region is >99.96% covered by the sequence data. (A) SNP and gene density throughout the target MHC region. Blue line: The number of dbSNP markers in discrete windows of 10 kbp. Orange diamonds: Genes that are involved in immunity. Black diamonds: Other genes. Red text: Classical HLA class I and II genes. Areas of significant structural diversity are indicated with green brackets; these are HLA-DR and C4A/C4B. Build 142 of dbSNP was used, which has 225,302 SNP sites mapped to the target region. The full list of genes in the target region is shown in Supplemental Table S3A. (B) Depth of sequence reads (duplicates removed) derived from the COX cell line following stringent alignment to the COX reference MHC haplotype. (C) Read depth after sequence reads derived from chimpanzee Clint had been mapped to the reference sequence (also derived from Clint) and PCR duplicates removed. Stringent alignment criteria were used to map the sequence reads back to the chimpanzee genomic segment that is equivalent to the human MHC region (the panTro4 MHC, Chr 6: 28774516-33956232).











