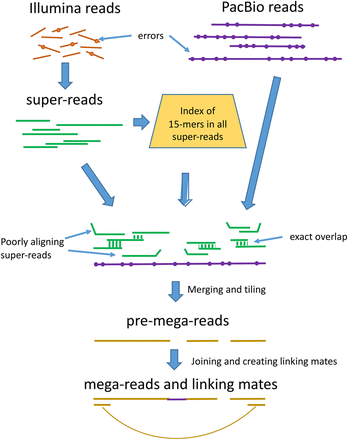
Overview of the mega-reads algorithm. Low-error rate Illumina reads (top left) are used to build longer super-reads (green lines), which in turn are used to construct a database of all 15-mers in those reads. PacBio reads (purple lines) and super-reads are then aligned, using the 15-mer index. Inconsistent super-reads are shown as kinked lines; these are discarded, and the remaining super-reads are merged, using the PacBio read as a template, to produce pre-mega-reads (yellow). These are further merged to produce the final mega-reads and to generate linking mates across gaps.











