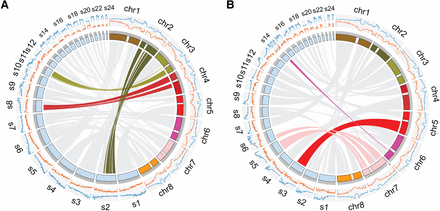
Comparing the assemblies of E. syriacum and C. planisiliqua to the ancestral karyotype present in the genome of A. lyrata. The eight chromosomes of A. lyrata are shown in colored blocks. Centromeric regions are indicated by white breaks. Scaffolds of the assemblies generated here of more than 1 Mb are shown in light blue blocks. The two histograms outside of chromosome karyotypes show the gene (orange) and repeat (blue) densities assessed with window sizes of 1 Mb for A. lyrata and 200 kb for E. syriacum or C. planisiliqua. (A) Three scaffolds of E. syriacum include similarities to the two flanking regions of A. lyrata CEN2, CEN3, and CEN4. (B) Scaffolds 3, 5, 6, and 14 include up to 7 Mb of putative centromeric regions, which are absent in the core assembly of A. lyrata, as these regions do not show any homology to any region in the A. lyrata genome.











