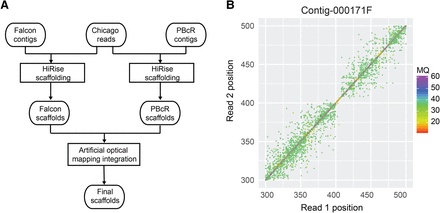
Assembly scaffolding using chromosome conformation capture data. (A) Improved chromosome conformation capture data scaffolding workflow. (B) Misassembly identification using chromosome conformation capture read pairs. The paired-end mapping positions in the region 300–500 kb of FALCON contig 000171F show a sudden absence of read pairs spanning across the region at around 410 kb. A misassembly at this region was indicated by HiRise. (MQ) Mapping quality.











