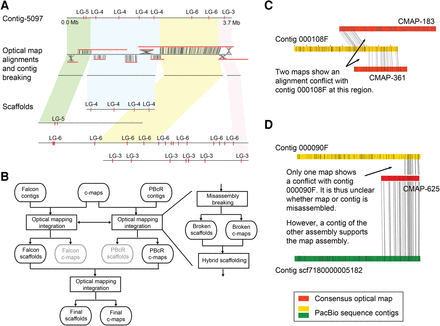
Optical mapping-based assembly correction and scaffolding. (A) Example of misassembly breakage and new scaffolding using optical mapping data. Three misassemblies in contig-5097 were identified with the optical map alignments (and also validated by the genetic maps; markers shown with red ticks). The original contig was broken, and the subsequent scaffolding of the four contigs, which resulted from breaking the original contig at the misassemblies, introduced them into the context of larger scaffolds, which were supported by the genetic map. (LG) Linkage group. (B) Improved optical mapping scaffolding workflow. Integration of optical mapping information includes breakage of misassembled contigs and consensus maps (c-maps) followed by hybrid scaffolding. (C) FALCON contig 000108F is apparently misassembled, as two different consensus maps (CMAP-183 and CMAP-361) have conflicting alignments with the same region of this contig. (D) A conflict between FALCON contig 000090F and CMAP-625 is not sufficient to decide on the origin of the underlying misassembly. However, CMAP-625 can be fully aligned to contig scf7180000005182 of a different (PBcR) assembly, supporting the correctness of this consensus map and thereby suggesting a misassembly in the contig.











