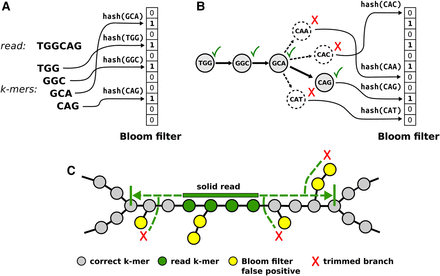
Overview of the ABySS 2.0 assembly algorithm. (A) k-mers from each input sequencing read are loaded into the Bloom filter by computing the hash values of each k-mer sequence and setting the corresponding bit in the Bloom filter. For clarity, we show a Bloom filter that uses a single hash function; in practice, multiple bit positions are set for each k-mer using multiple independent hash functions. (B) A path in the de Bruijn graph is traversed by repeatedly querying for possible successor k-mers and advancing to the successor(s) that are found in the Bloom filter. Each possible successor corresponds to single-base extension of the current k-mer by “A,” “C,” “G,” or “T.” (C) ABySS 2.0 builds unitig sequences by extending solid reads left and right within the de Bruijn graph. A solid read is a read that consists entirely of k-mers with an occurrence count greater or equal to a user-specified threshold (solid k-mers); the optimum minimum occurrence threshold is typically in the range of two to four. Extension of a solid read is halted when either a branching point or a dead end in the de Bruijn graph is encountered. A look-ahead algorithm is employed to detect and ignore short branches caused by Bloom filter false positives and/or recurrent read errors.











