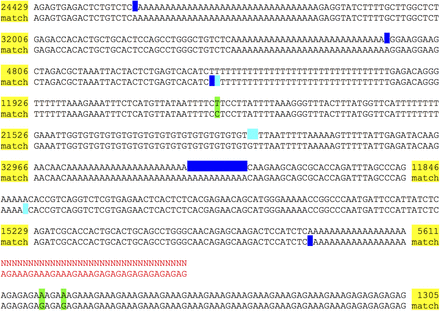
Alignment of Supernova assembly to finished sequence from the same sample. GenBank sequence AC004551.1 for finished clone RPCI1-71H24 has length 162,346 bases, and its reverse complement perfectly matches GRCh37. The clone encompasses a region of Neandertal origin (Mendez et al. 2013). Both the clone and assembly F (Table 1) represent DNA from the same HGP donor. The clone matches a region of which 96% is between two megabubbles in the assembly, thus represented as homozygous. The alignment of the assembly to the clone region on GRCh37 is shown. Each line pair shows the assembly on top and the reference on the bottom. (Yellow) abbreviated, perfectly matching stretches; (green) mismatched bases; (blue) indels; (cyan) indels, but not present in comparison to raw graph; (red) captured gap: signified by 34 Ns (actual number in assembly is 100); assembly region also has two cycles, each suffixed by 10 Ns in output, not shown. In these cases the flattened sequence for the cycle exactly matches the reference.











