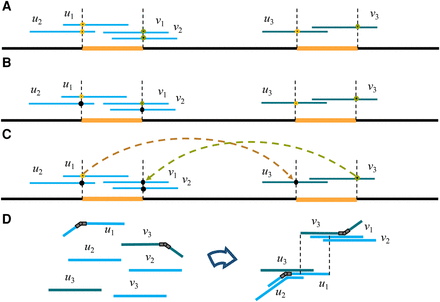
The Contagion algorithm. (A) Two unbridged repeats are shown as orange segments. (B) The Contagion algorithm first kills the start-repeat annotation at u2 (due to its overlap with u1) and the end-repeat annotation at v2 (due to its overlap with v1). (C) The Contagion algorithm then kills the start-repeat annotation on u3 (due to its internal match with u1) and the end-repeat annotation on v1 (due to its internal match with v3). (D) Finally, an in-hinge is placed on u1 and an out-hinge is placed on v3. During the hinge-aided greedy assembly step, hinges allow u2 and u3 to choose the in-hinge at u1 as their successor. Similarly, v1 and v2 pick the match starting at the out-hinge on v3 as their predecessor match.











