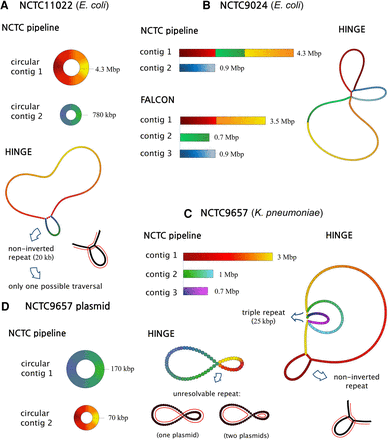
Analysis of HINGE graphs on selected data sets. By identifying unbridged repeats, collapsing them, and then performing resolutions based on uniquely traversable loops, HINGE prevents misassemblies and produces a user-friendly, interpretable assembly graph. We color the graph nodes according to their corresponding position on the NCTC pipeline contigs. (A) On NCTC11022, HINGE identifies an unbridged repeat, which is later resolved. (B) On NCTC9024, HINGE identifies an unbridged triple repeat (with one inverted copy), which cannot be resolved due to the existence of three distinct traversals of the graph. (C) HINGE identifies an unbridged triple repeat. (D) HINGE identifies an unresolvable repeat shared by two small plasmids.











