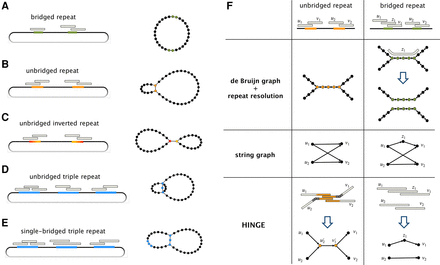
The goal of HINGE is to produce a maximally resolved assembly graph, where repeats that are bridged by the reads are not collapsed, and repeats that are unbridged are collapsed in a natural way, similar to what is achieved with de Bruijn graphs. (A) If at least one of the two copies of a repeat is bridged (green segments), the maximally resolved assembly graph should separate the two copies. In (B–E), respectively, we illustrate an unbridged repeat, an unbridged inverted (i.e., reverse-complemented) repeat, an unbridged triple repeat, and a single-bridged triple repeat, and the assembly graph obtained by collapsing segments corresponding to unbridged repeats. Notice that in B,E, the graph admits a single traversal and can be further resolved, while in C,D, the graph admits two distinct traversals and cannot be further resolved (see Supplemental Fig. S15). (F) The representation of a bridged and an unbridged repeat in the de Bruijn graph approach, in the standard string graph approach and according to HINGE. The de Bruijn graph approach collapses the repeated segment, which allows a natural repeat resolution step if a bridging read is found. The representation in the string graph (if there is no read entirely contained in the repeat) is an hourglass-like motif. HINGE emulates the de Bruijn graph layout but in an overlap graph framework.











