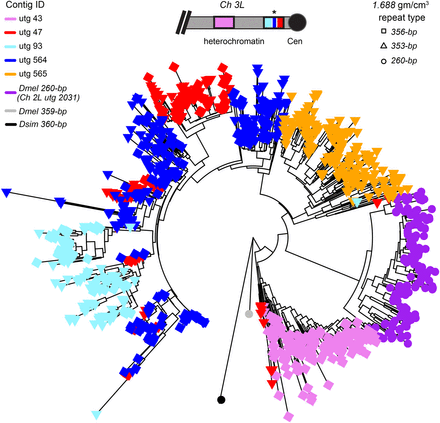
Neighbor-joining tree of D. melanogaster 1.688 family satellites: the 353/356-bp satellites on Chromosome 3L, the 260-bp locus on Chromosome 2L and a single consensus repeat of 359-bp from the X Chromosome, and a related repeat from D. simulans (360-bp). Tips are colored according to the contig from which they originate. The tip symbol refers to monomer repeat type: 353-bp, 356-bp, or 260-bp. The inset shows a schematic of the Chromosome 3L 1.688 loci and the organization of their respective contigs (the two clusters are ∼2 Mb apart). Contig utg 564 does not align to the reference genome, but we infer its location (*) based on repeats clustering with those in utg 47 and a gap in the reference at this genomic location. Contig utg 565 is unmapped.











