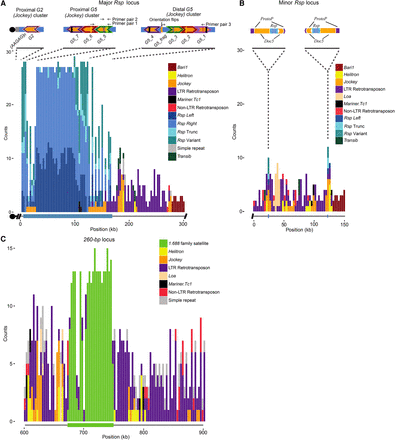
Maps of complex satDNAs contigs. Counts for each repetitive element family in our custom Repbase library were plotted in 3-kb windows across each contig. (A) Rsp locus on Chromosome 2R. Blue bars correspond to Left, Right, variant or truncated repeats, whereas other colors correspond to various TE families as indicated to the right of each contig. Rsp spans ∼170 kb of the 300-kb contig (thick blue line below the x-axis). Above the plot is a schematic showing the orientation of two G5 clusters flanking the Rsp locus and a separate contig containing Rsp and the Jockey element G2, which is directly adjacent to AAGAG satellite repeats. The colors of the chevron outlines indicate the G5 elements with the highest degree of similarity with one another. Solid and dashed lines within the insertions show the approximate locations of shared insertions or deletions, respectively. Several configurations of indels are unique, such as the two in G5_5 or the deletion in G5_1, which allows verification of the cluster. The G2 contig may contain the most centromere-proximal repeats (black circle; see text). (B) Minor Rsp locus on Chromosome 2R. The inset shows the detailed orientation of the two clusters (five Rsp repeats per cluster, ∼100 kb apart); the direction of arrows indicates the relative orientation of the elements. The Rsp repeats (blue chevrons) are nested within Doc5 (orange chevrons) insertions, which are in turn nested within insertions of a transposon known as ProtoP (purple chevrons). The clusters of Rsp+Doc5+ProtoP share ∼96% sequence identity with one another, and are in an inverted orientation. (C) 260-bp locus on Chromosome 2L. Only the area surrounding the 260-bp array is shown (300 kb of ∼1.1-Mb contig). The 260-bp locus spans ∼70 kb of the 1.1-Mb contig (green below the x-axis) and is interrupted with Copia transposable elements.











