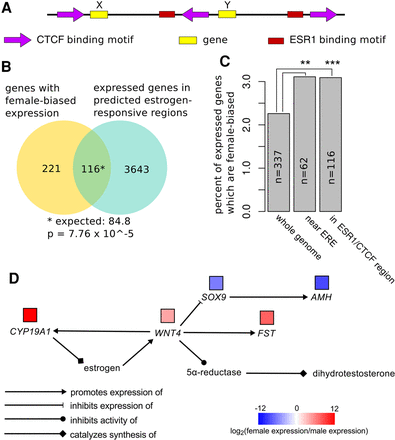
Genes in regions of the genome predicted to be under estrogenic regulation of gene expression are significantly more likely to be female biased in the post-TSP gonads. (A) Our model for predicting regions of the genome under estrogenic regulation of gene expression, based on the CTCF extrusion model (Sanborn et al. 2015) and the Chan and Song model of estrogen receptor binding site activity (Chan and Song 2008). In this example, Gene X is predicted to be estrogen responsive and Gene Y is not because Gene X is between two inward-oriented CTCF binding motifs along with an ESR1 binding site, while Gene Y is not. (B) Of the 14,943 genes expressed in the post-TSP gonads, 337 have female-biased expression and 3759 are in predicted estrogen-responsive genomic regions. However, 116 of these genes are both female biased and within predicted estrogen-responsive regions, a significantly higher number than the expected 84 (P = 7.76 × 10−5). (C) Percentages of expressed genes with female-biased expression in the whole genome versus near an estrogen response element and in a predicted estrogen-responsive CTCF region. Regions near an estrogen response element and predicted estrogen-responsive regions are both enriched for female-biased genes. (**) P ≤ 0.01; (***) P ≤ 10−4. (D) Pathway diagram showing results of increased CYP19A1 expression after the TSP in the gonads of embryos incubated at FPT. Sex-bias fold-changes for each gene in the pathway are shown in boxes above the genes.











