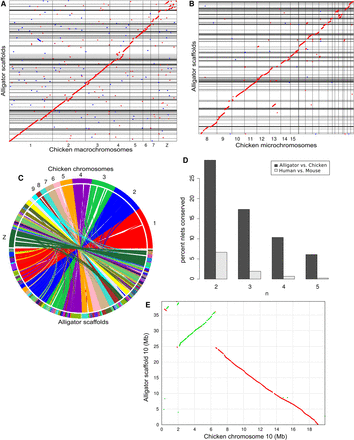
Our new long-range assembly of the American alligator genome allows analysis of the synteny between crocodilians and birds. (A,B) Dot plots of an anchored whole-genome alignment between the chicken and American alligator genomes show a high degree of synteny, with many long alligator scaffolds covering significant portions of chicken chromosomes, including macrochromosomes (A) and microchromosomes (B). (C) A circle plot of synteny between the alligator and chicken genomes made using SyMAP (Soderlund et al. 2011). (D) Conservation of ordered gene doublets, triplets, quadruplets, and quintuplets between alligators and chickens versus between humans and mice, showing much higher synteny between alligators and chickens than between humans and mice. (E) Alligator scaffold 10 covers a vast majority of the chicken microchromosome 10. However, there are several small inversions and one large inversion between the two. Green and red dots represent forward and reverse matches, respectively.











