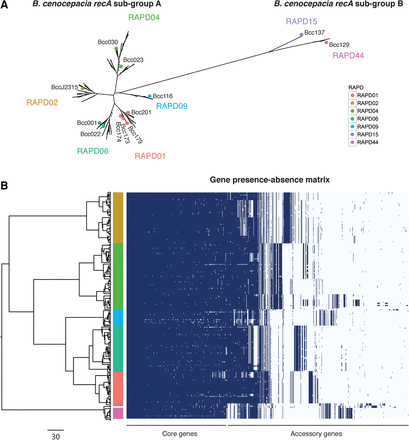
Core gene phylogeny and gene presence across B. cenocepacia isolates. (A) Core genome phylogeny built using RAxML (Stamatakis 2014) from 2148 orthologous clusters of protein coding genes from 209 B. cenocepacia isolates (four NCBI references were included, and 10 Illumina assemblies with more than 500 contigs were excluded). The structure shows that the RAPD genotypes form monophyletic clades. Colored dots correspond to RAPD reference genomes sequenced using PacBio and represent at least one isolate from each epidemic lineage. (B) Left side shows the same phylogenetic tree based on a concatenated core gene alignment. The right side shows a gene possession matrix, with each row representing a strain's gene content. Each column corresponds to a homologous gene cluster (i.e., after merging “orthologous” clusters into homologous clusters), and columns are ordered by the frequency of gene presence. The results clearly indicate extensive clade-specific gene content. The colored bar indicates RAPD genotype. Scale bar for the phylogenetic tree on the bottom left represents the number of SNPs in core genes.











