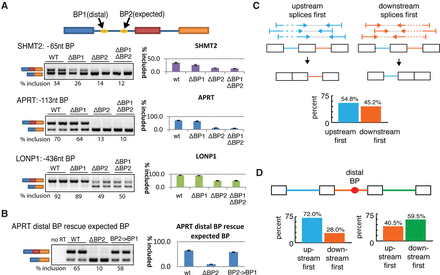
Distal branchpoints are functional elements that are associated with delayed splicing. (A) Both or either one of the distal (BP1) and expected (BP2) branchpoints was mutated in a splicing minigene as depicted at the top. Minigenes were transfected into HEK293T cells and inclusion of the middle exon was measured by RT-PCR as a readout of branchpoint activity (two replicates displayed for each construct). The splicing inclusion level was estimated by ImageQuant software across ≥3 replicates and recorded below each panel and in the histogram to the right. (B) Expected branch site of APRT (i.e., BP2) was replaced by the sequence of the distal branch site (BP2→BP1) and the functionality of branch sites was assayed by RT-PCR as in A. (C) Scheme for determining relative order of intron removal from paired-end reads, using all human introns. The bar plot indicates the degree to which the upstream (blue) and downstream (orange) introns splice first. (D) Introns that contain distal branch sites (orange) were analyzed for order of intron removal. “Upstream first” contains loci where the upstream intron reliably spliced first (i.e., >90%) and “downstream first” contains loci where the distal intron spliced before the upstream intron (left). Similar comparisons were performed on the downstream intron (right).











