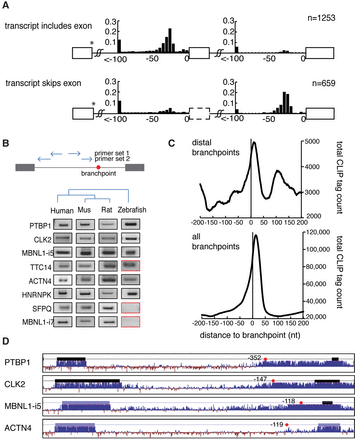
Distal branchpoints are conserved and associated with alternative exon usage. (A) The location of branchpoints was mapped in exon inclusion (top) and exon skipping (bottom) transcripts. Distal branchpoints more than 100 nt upstream of the 3′ss were binned at −100 nt. y-axes indicate the fraction of the bars. All branchpoints depicted use the same donor site (marked by *). (B) Distal branchpoints were validated by nested lariat PCR in human, mouse, rat, and zebrafish. Sequencing was used to distinguish conserved branch site usage (black box) from nonconserved (red). (C) The CLIP signal of U2AF2 binding was mapped relative to distal branchpoints (top) and all branchpoints (bottom). (D) Conservation of four introns that contain distal branchpoints. Thick bars indicate exons, and the red dots are distal branchpoints.











