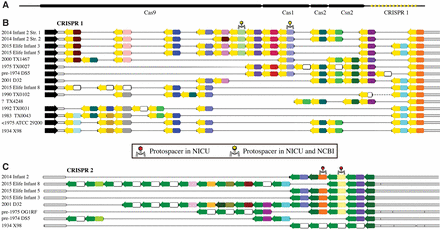
CRISPR spacers are maintained over decades in Enterococcus faecalis. (A) Genomic organization of CRISPR-Cas array #1. (B) Alignments of array #1 and (C) array #2 from E. faecalis from Infant 2 of this study compared to arrays reconstructed from publicly available genomes for isolates. The year of isolation of all E. faecalis isolates is provided to the extent possible. Infants marked “Elife” are those from a previous publication from the same NICU (Raveh-Sadka et al. 2015). Arrows represent repeats and colors represent spacers; identical colors symbolize identical spacers, whereas white spacers are unique. Phage symbols represent spacers with a protospacer match (max 1 mismatch) in a sequence assembled from infants in the same NICU as this study (red), and spacers with matches in both the same NICU and a separate genome in NCBI (yellow).











