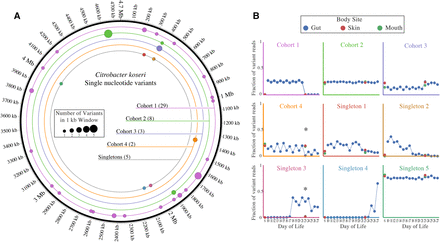
Subpopulations exist within colonizing C. koseri populations. (A) Single nucleotide variants were identified by mapping reads from all Infant 1 samples to the draft genome of C. koseri recovered from Infant 1. The total number of variants in each cohort is listed in parentheses. (B) The frequency of each variant in each sample. Cohorts are plotted as the average of all variant frequencies, with error bars representing standard deviation of the mean. Asterisks represent cases where the frequency of variants is statistically different between body sites (Fisher's exact t-test with Bonferroni correction, P < 0.01).











