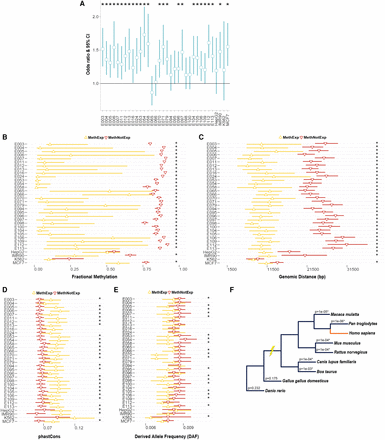
Association of distal CGI with the expression of MethExp genes. (A) Odds ratio and 95% confidence interval (CI) (y-axis) of the proportion of unmethylated CGIs upstream of MethExp genes versus MethNotExp genes in 34 tissue types (x-axis). A depletion of methylation at CGIs upstream of MethExp genes corresponds to a higher odds ratio. (B–E) Comparison of various properties for the CGIs upstream of MethExp genes (yellow) versus MethNotExp (red) genes; (B) fractional methylation level, (C) genomic distance to gene, (D) phastCons scores, and (E) derived allele frequencies (DAFs). The median and the 95% CI (x-axes) are shown for 34 tissue types (y-axes). (F) Phylogenetic tree of the eight vertebrate species used in determining the extent of shared synteny with human among methylated-promoter genes. The association between CGI-gene synteny and whether the gene is MethExp or MethNotExp was assessed via regression, while controlling for genomic distance between CGI and the gene. The significance of the association (P-value) is shown on each branch corresponding to the species used to estimate synteny with respect to human. Statistically significant associations (P < 0.05) are annotated with an asterisk in all plots.











