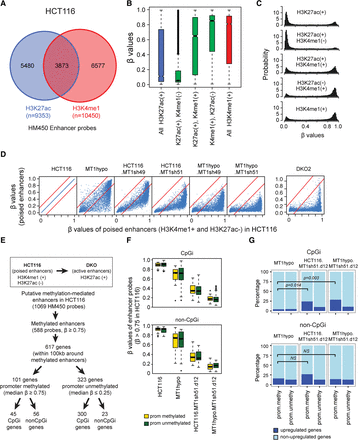
DNA methylation profiles in gene enhancers. (A) Infinium Methylation 450K (HM450) probes that overlap with putative enhancers marked by either H3K27ac or H3K4me1. (B,C) DNA methylation levels of putative enhancers in HCT116 depicted in whisker box plots (B) and histograms (C). In B, β values (y-axis) are matched to enhancer histone modification status as determined by ENCODE (encodeproject.org) (The ENCODE Project Consortium 2012) (x-axis). (D) DNA methylation levels of poised enhancers (H3K4me1+ and H3K27ac−) in HCT116-derived clones with various DNMT1 protein levels. The x-axis shows β values for wild-type HCT116 cells (range; first panel); the y-axis shows β values for DNMT1 knockdowns or knockouts and DKO cells. (E) Flow chart for selection of putative enhancers stratified by methylation analyses and that have been denoted by ENCODE to be poised in HCT116 wild-type cells and that became active in HCT116-DKO cells (Blattler et al. 2014). The putatively controlled genes for these enhancers (50 kb upstream or downstream) have been determined by informatics analyses outlined in the text. HM450 probes that are located within the poised enhancers and heavily methylated (β values ≥0.75) are included in the analyses. (F) Whisker box plots for methylation levels (y-axis) of poised enhancer probes (β values ≥0.75) in HCT116 cells with different DNMT1 levels. Genes are grouped according to their promoter CpG islands and methylation status. The yellow box represents genes with methylated promoters (average β values ≥0.75), and the green box represents genes with unmethylated promoters (average β values ≤0.25). (G) Percentage of genes (y-axis) with up-regulated (defined by being 1.4-fold or greater than basal values in wild-type HCT116 cells; dark blue portion of bar) versus unchanged expression (light blue portion of bar) following the DNMT1 depletions outlined in F (shown at the tops of the panels). The analysis included 617 genes defined in E. P-values, indicated by the bars in the panels, are calculated for Fisher's exact test.











