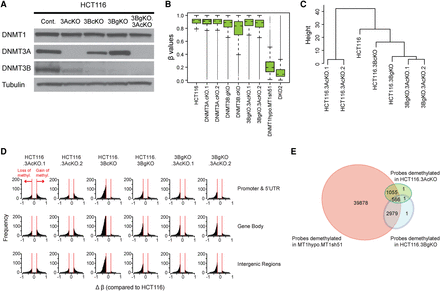
Effects of depleting DNMTs 3A and 3B on DNA methylation maintenance. (A) Western blot analysis of DNMTs in HCT116 cells subject to depletion of DNMT3s using CRISPR (3AcKO, 3BcKO) or genetic knockout approach (3BgKO). DNMT3 double knockout cells are created by CRISPR knockout of DNMT3A in 3BgKO cells (3BgKO.3AcKO). (B) Whisker box plots of DNA methylation levels assayed by Infinium 450K array in HCT116 and the derivative cell lines depleted of individual DNMTs (x-axis). All probes with β values greater than 0.75 were included in the analyses. (C) Hierarchical cluster analysis of DNA methylation patterns in HCT116 WT and DNMT mutant cells. Euclidean distance matrixes were used for the clustering based on complete linkage agglomerative algorithm. The top 10,000 most variable probes were included in the analysis. (D) Histograms of frequency distribution (y-axis) of differentially methylated (Δβ ≥ 0.2) Infinium 450K CpGi probes (x-axis) in DNMT3 knockout cells compared to the HCT116 parental cells. (E) Venn diagram of Infinium 450K probes demethylated by DNMT1 shRNA knockdown (MT1hypo.MT1sh51), DNMT3A CRISPR knockout (common between HCT116.3AcKO.1 and HCT116.3AcKO.2), and DNMT3B genetic knockout (HCT116.3BgKO). The analysis includes promoter probes with β values greater than or equal to 0.75 in parental HCT116 cells and showing a decrease by more than 0.2 upon depletion of individual DNMTs.











