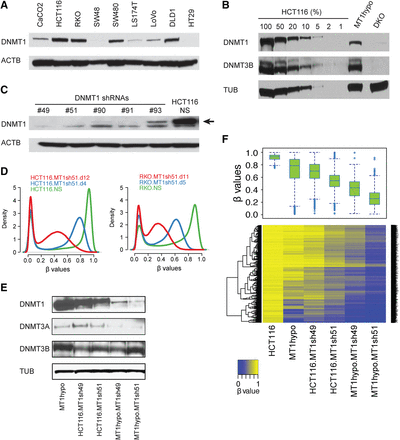
Relationships of DNMT1 levels to the maintenance of DNA methylation in colon cancer cells. (A) Western blot analysis of DNMT1 in a panel of colorectal cancer cell lines: (ACTB) beta actin. (B) Western blot analysis of DNMT1 and DNMT3B from HCT116 DNMT1 hypomorph cells (MT1hypo) and compared to dilutions of extracts of wild-type HCT116 cells: (TUB) alpha tubulin; (DKO) HCT116 double knockout cells (carrying a DNMT1 hypomorph and DNMT3B knockout). (C) Western blot analysis of DNMT1 in HCT116 cells subject to DNMT1 shRNA knockdowns: (NS) nonsilencing shRNA control. An arrow denotes the DNMT1-specific band. (D) Genome-wide methylation profiles measured by Illumina Infinium HumanMethylation450K array in HCT116 and RKO subjected to DNMT1 shRNA knockdown (MT1sh51) for different lengths of time. The density plots include all probes on individual arrays for each sample. The x-axis indicates β values: scores in the range between 0 and 1 indicating the level of DNA methylation (lowest to highest = increasing DNA methylation). The y-axis indicates the probability densities which describe the distribution of β values for all probes. (E) Western blot analysis of DNMTs in MT1hypo and HCT116 cells subject to DNMT1 knockdowns with different shRNAs. (F) DNA methylation analyses by Illumina Infinium HumanMethylation 450K array in HCT116 cells with decreasing DNMT1 levels shown in E. A whisker box plot of the resultant β values is shown (upper) with a heatmap of the β values (lower). Note the step-wise reduction of DNMT1 that correlates with progressive demethylation. The analysis includes methylated promoter CpGi (CpG island) probes with β values greater than or equal to 0.75 in wild-type HCT116 cells.











