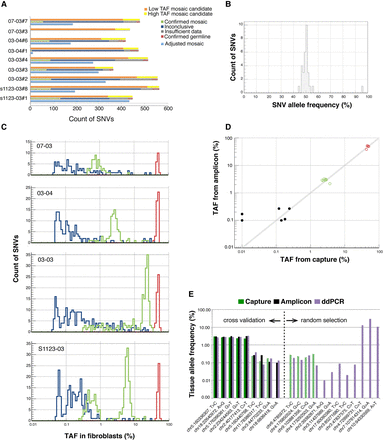
Discovery and confirmation of LM-SNVs in children. (A) Discovered LM-SNVs in hiPSC lines were divided into two groups: low tissue allele frequency (with no evidence in fibroblasts, orange bars), and high TAF (with some evidence, yellow bars). Site reanalysis in fibroblasts with DNA capture and deep sequencing confirmed that, on average, 74 LM-SNVs in each hiPSC line are mosaic SNVs in fibroblasts (green bars) or 235 when adjusted for discovery sensitivity and ascertained fraction (light blue bars). (B) Virtually all LM-SNVs were present at around 50% AF in hiPSC lines as detected by amplicon-seq experiments. (C) Capture-seq experiment in fibroblasts revealed that mosaic SNVs were present in the fibroblast tissue with TAF ranging from 0.25% to 35%. Distributions of TAF have clear peaks. (D) Amplicon-seq experiment for 57 LM-SNVs sites results in better sensitivity and confirms an additional six LM-SNVs with low TAF as mosaic (black dots), including two with no supporting read in the data from capture (shown with TAF of 10−4 for capture). Germline and confirmed mosaic SNVs by capture experiments are in red and green circles, respectively. (E) ddPCR reactions revealed excellent concordance in TAF estimates with the capture-seq and amplicon-seq experiments. Dashed green bars show SNV sites for which capture experiments were conducted, but support for the alternative allele was consistent with background sequencing noise. Additional ddPCR assays confirmed mosaic SNV at even lower TAFs that could not be accessed by the other two experiments.











