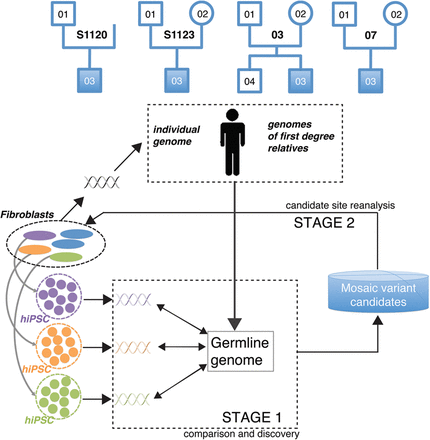
Conceptual diagram of our approach. Our cohort consisted of four families, each having a proband with autism, while the other family members were phenotypically normal. Family 03 includes a normal male sibling. Three hiPSC lines were generated from the fibroblast samples of each person in the cohort. As hiPSC lines are clonally derived from single cells, comparison (STAGE 1) of their genomes to the germline genome uncovers mosaic variants present in the founder fibroblast cells of hiPSC colonies (green, orange, and purple variants). Germline variants for children were inferred from corresponding parents and those for parents from corresponding children and fibroblast samples. Analysis at STAGE 1 yields a list of putative mosaic variants manifested in hiPSC lines. In STAGE 2, the mosaic candidates are scrutinized by additional experiments in founder fibroblasts to confirm their presence and to determine their tissue allele frequency (TAF). Naming pattern for hiPSC lines is as follows: family-person#hiPSC, e.g., S1120-01#2.











