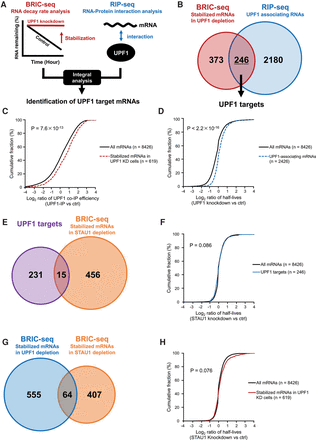
Identification of UPF1 and SMD target mRNAs, including UPF1 targets partially regulated by the STAU1 protein. (A) Schematic of the method to identify UPF1 targets based on the combinatorial analysis of BRIC-seq and RIP-seq. (B) Venn diagram illustrating the overlap of the two types of mRNAs: those with more than twofold prolonged half-lives upon UPF1-knockdown in HeLa tet-off cells (red: BRIC-seq, 619 mRNAs) and those enriched by more than twofold in the HA-UPF1 immunoprecipitation fraction relative to input fraction (blue: RIP-seq, 2426 mRNAs). The intersection of stabilized mRNAs in UPF1-knockdown cells (BRIC-seq) and UPF1-associating mRNAs (RIP-seq) were judged as bona fide UPF1 target mRNAs (246 mRNAs). (C) Cumulative distribution of changes in UPF1 association of 619 transcripts stabilized upon UPF1 depletion (red dashed line; median: 0.66) and 8426 whole transcripts (black solid line; median: 0.17). Changes in UPF1 association were calculated as the number of mapped tags of RNA-seq immunoprecipitated with HA-UPF1 by reads per kilobase of transcript per million mapped reads [RPKM]/the number of mapped tags of input RNA-seq [RPKM] ≥2. P-values were calculated using the Wilcoxon rank-sum test. (D) Cumulative distribution of changes in RNA stability of 2426 UPF1 associating transcripts (blue dashed line; median: 0.02) and 8426 whole transcripts (black solid line; median: −0.20). Changes in RNA stability were determined by the following calculation: the half-life of each transcript in UPF1 depletion/the half-life of each transcript in control. P-values were calculated using the Wilcoxon rank-sum test. (E) Venn diagram illustrating the overlap of two types of genes: UPF1 target mRNAs (purple: 246 genes) and mRNAs with more than twofold prolonged half-lives upon STAU1-knockdown in HeLa tet-off cells (orange: BRIC-seq, 471 genes). The intersection of the UPF1 target mRNAs and stabilized mRNAs in STAU1-knockdown cells (BRIC-seq) indicates STAU1-mediated mRNA decay (SMD) target mRNAs (15 genes). (F) Cumulative distribution of changes in RNA stability of 246 UPF1 target transcripts (blue line; median: 0.032) and 8426 whole transcripts (black line; median: 0.000). Changes in RNA stability were determined by the following calculation: the half-life of each transcript in STAU1 depletion/the half-life of each transcript in control. P-values were calculated using the Wilcoxon rank-sum test. (G) Venn diagram illustrating the overlap of two types of mRNAs: those with more than twofold prolonged half-lives upon UPF1-knockdown (blue: BRIC-seq, 619 genes containing 246 UPF1 targets and 373 transcripts stabilized by UPF1 depletion but not associated with UPF1; see B) and STAU1-knockdown in HeLa tet-off cells (orange: BRIC-seq, 471 genes). The number of the stabilized mRNAs upon UPF1-knockdown (BRIC-seq) and those upon STAU1-knockdown (BRIC-seq) contains 64 mRNAs. (H) Cumulative distribution of changes in RNA stability of mRNAs with more than twofold prolonged half-lives upon UPF1-knockdown (red line; median: 0.008) and 8426 whole transcripts (black line; median: 0.000). Changes in RNA stability were determined by the following calculation: the half-life of each transcript in STAU1 depletion/the half-life of each transcript in control. P-values were calculated using the Wilcoxon rank-sum test.











