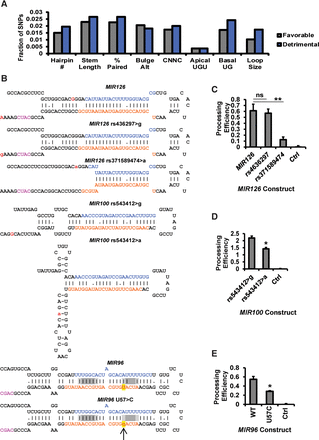
The effects of human SNPs on pri-miRNA processing. (A) Human SNPs in dbSNP human Build 142 that are located close to human pri-miRNAs were evaluated for their impact on hairpin structure and sequence features. For each SNP, hairpin features for minor and major alleles were compared. The fractions of minor alleles that have predicted favorable (gray) or detrimental (blue) impact (relative to the major allele) on pri-miRNA processing were plotted. (B) Predicted hairpin structures of major and minor alleles tested in C through E. Color-coded elements include 5p mature miRNA (blue), 3p miRNA (orange), major allele base (red letter; uppercase), and minor allele base (red letter; lowercase). Watson-Crick pairings are indicated with vertical bars, whereas G:U pairings are indicated with dots. The positions of the bulge-depleted regions, as measured from the base of the stem, are shaded in gray. Arrow and yellow-highlighted red text indicate the sequence variation. (C–E) The indicated pri-miRNA constructs were subjected to processing reporter assay in BaF3 cells. Processing efficiencies were normalized to mouse Mir125b-2 WT (set to one) and an empty vector control (Ctrl; set to zero). N = 3. Error bars, SD. (*) P < 0.05; (**) P < 0.01; (ns) not significant.











