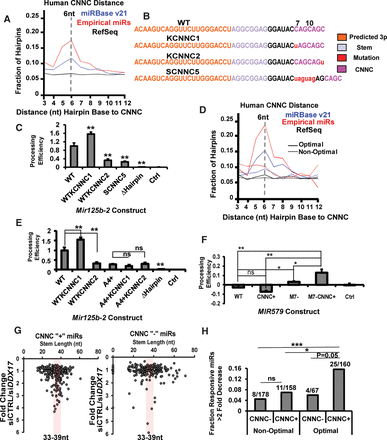
The roles of the CNNC motif in pri-miRNA processing. (A) The distributions of putative CNNC motifs were plotted for human hairpins from miRBase, Empirical miRNA set, or RefSeq. The distance of CNNC motif was measured in nucleotides from the base of the stem to the first C in the motif. The most enriched position (6 nt) was highlighted. (B,C) Processing of WT mouse Mir125b-2 or its CNNC mutants was assayed. The sequences of CNNC mutants were depicted in B. Color-coded elements include the 3p miRNA shown in orange, nucleotides in the stem shown in lavender, mutated or inserted nucleotides shown in red, putative CNNC motifs in magenta, and positions relative to stem base indicated by numbers. (C) BaF3 cells were transduced with the processing reporters, and the processing efficiencies were measured, with the level for WT mouse Mir125b-2 set to one and the level of an empty vector (Ctrl) set to zero. A construct removing the hairpin in mouse Mir125b-2 (ΔHairpin) was also used as a control. N = 3. P-values were annotated for comparison with WT construct. (D) Distributions of the CNNC motifs, among miRNA or non-miRNA hairpins that were of optimal (33–39 nt) or nonoptimal (<33 or >39 nt) length. CNNC motifs preferentially co-occur with optimal length miRNA hairpins. (E) WT mouse Mir125b-2 or combination mutants containing stem length alterations (A4+) (Fig. 2) and CNNC mutations (see C) were measured in processing reporter assays in BaF3 cells. N = 3. (F) Processing efficiencies for WT MIR579, which is both long (44 nt of stem length) and without a CNNC motif, and its mutants were measured in BaF3 cells. Mutant structures are illustrated in Supplemental Figure S4K, with the CNNC+ mutant containing an engineered putative CNNC motif 8 nt from stem base of MIR579, the M7-mutant shortening the stem length by 7 nt, and the M7-CNNC+ mutant having both shortened stem length and a CNNC motif. Data were normalized the same way as in B. N = 3. (G) The fold-decrease of mature miRNA expression upon siRNA knockdown of DDX17 versus a control siRNA (siCtrl) was plotted against the stem length. Each dot represents a single miRNA, with those containing putative CNNC motifs (CNNC+) and without (CNNC−) plotted separately. The optimal stem length range (33–39 nt) was highlighted. (H) Data from G were plotted to quantify the number of miRNAs with decreased expression (greater than twofold) upon DDX17 knockdown, for miRNAs with optimal or nonoptimal stem length, and with putative CNNC or without. Numbers above the bars indicate the number of decreased miRNAs out of all miRNAs in the indicated category. Error bars, SD. (*) P < 0.05; (**) P < 0.01; (***) P < 0.001; (ns) not significant.











