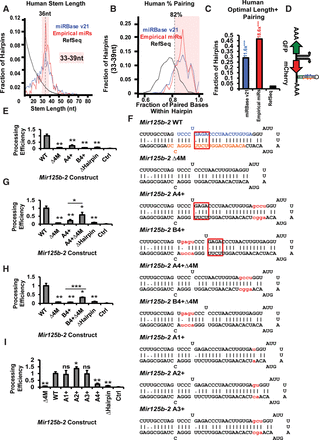
Optimal stem length is required for efficient processing. (A) The distributions of the stem lengths of human hairpins were plotted for pri-miRNAs in miRBase, Empirical pri-miRNAs, and RefSeq hairpins. The optimal range of 36 ± 3 nt is highlighted. (B) The distributions of percentage pairing within hairpin stem were plotted for human hairpins. A cutoff of 82% pairing was highlighted. (C) The fractions of human hairpins with both optimal stem length (33–39 nt) and pairing (≥82%) were plotted, with fold enrichment over RefSeq indicated. (D) Diagram of the lentiviral pri-miRNA processing reporter. (E–I) The processing of wild-type (WT) and mutant mouse Mir125b-2 was measured in BaF3 cells. Data were normalized with the processing level for WT mouse Mir125b-2 construct set to one and the level of an empty vector (Ctrl) set to zero. A construct removing the hairpin in mouse Mir125b-2 (ΔHairpin) was also used as a control. N = 3. P-values are annotated above the bars for comparison with WT construct. Other P-value comparisons are indicated with horizontal bars. (F) Sequences and predicted structures of constructs tested in E and G through I. Color-coded elements include 5p mature miRNA (blue), 3p miRNA (orange), insertions (red letters; lowercase), and deletions (red box) that occurred in other related constructs. Watson-Crick pairings are indicated with vertical bars, whereas G:U pairings are indicated with dots. (*) P < 0.05; (**) P < 0.01; (***) P < 0.001; (ns) not significant.











