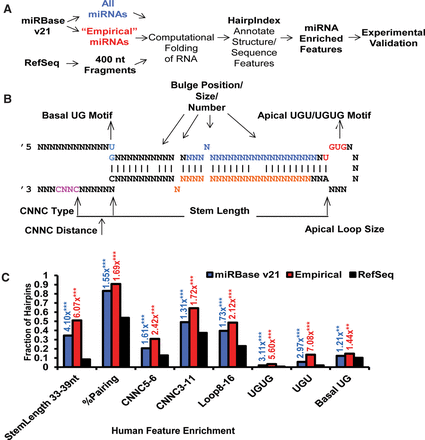
Systematic evaluation of miRNA and non-miRNA hairpins reveal enriched structural and sequence features in pri-miRNAs. (A) A schematic of the computational and experimental workflow. (B) Diagram of major hairpin features annotated by the HairpIndex pipeline. For pri-miRNA hairpins, blue letters represent 5p mature miRNA; orange letters, 3p mature miRNA. Primary sequence motifs—including CNNC, basal UG, and apical UGU/UGUG—were also highlighted. (C) The fractions of hairpins containing each indicated feature were plotted for human pri-miRNAs in miRBase v21, in a subset of “Empirical” miRNAs, or in human RefSeq fragments. The numbers above the bars indicate the fold enrichment of the corresponding feature over the RefSeq control. The numbers for stem length indicate the range of stem lengths in nucleotides; % pairing indicates the percentage of paired bases within the stem; the numbers after CNNC indicate the position range of putative CNNC motifs relative to the base of the hairpin, as measured in nucleotides; the numbers after loop indicate the range of the loop sizes; and basal UG, apical UGU, or apical UGUG motifs indicate the presence or absence of such features in hairpins. P-values were calculated using Fisher's exact test. (**) P < 0.005; (***) P < 0.0005.











