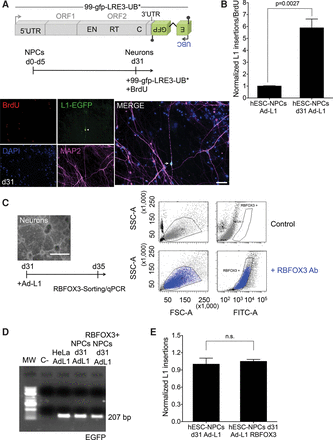
Efficient LINE-1 retrotransposition in differentiating NPCs. (A) Scheme of plasmid 99-gfp-LRE3-UB* and of DNA transfection-based experiments conducted in differentiating NPCs. (Below) Representative image of differentiating NPCs transfected at day 31 in the presence of BrdU and stained with L1-EGFP (green), MAP2 (pink), and BrdU (red); nuclear DNA was stained with DAPI (blue). The small pictures at the left side contain the independent captured images used in the merged picture. A white arrow marks a MAP2/L1-EGFP single cell that stains negative for BrdU. (White bar) 50 µm. (B) L1-EGFP copy number quantification in differentiating NPCs infected at the indicated time. The graph shows the normalized number of L1-EGFP sequences detected (L1 insertions) and has been corrected with infection efficiency values (using beta gal qPCR data) and for cell proliferation values. The P-value of the comparison (0.0027) and the SEM are also indicated. (C) Rationale of the RBFOX3 sorting-based assay. Differentiated NPCs (at day 31) were infected with the Ad-L1, and 5 d later RBFOX3-expressing cells were FACS-sorted and gDNA isolated. (Right) Representative FACS histogram plots of cells incubated (bottom) or not (top) with the anti-RBFOX3 antibody. (D) Results from the PCR-intron assay conducted on gDNAs isolated from Ad-L1 infected NPCs at day 31 and FACS-sorted (using a RBFOX3 antibody) 5 d after the infection. gDNA isolated from unsorted infected differentiated NPCs and infected HeLa cells were used as controls in these assays. (C- lane) PCR negative control without template. (E) L1-EGFP copy number quantification in differentiated Ad-L1 infected NPCs sorted for RBFOX3 expression. The graph shows normalized L1 insertions (i.e., the number of L1-EGFP sequences detected); for comparison, the value obtained in unsorted Ad-L1 infected hESC-NPCs at day d31 post-differentiation was designated 1. (n.s.) not significant. The SEM is also shown. In B and E, an unpaired Student's t-test was applied.











