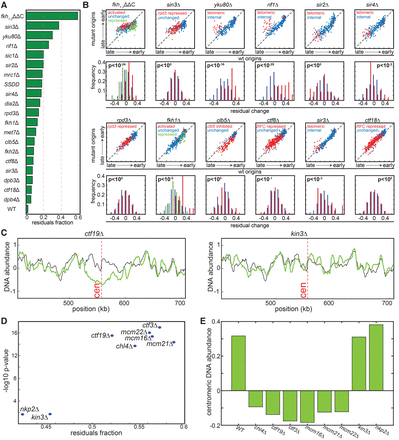
Mutants showing origin-specific effects. (A) Scoring mutants by the fraction of replication profile not explained by global changes. Shown are the fractions of the variances of the measured replication signals not explained by a model assuming no origin-specific effects. (B) Change in the replication timings of individual origins: For each mutant, the upper figure shows the inferred replication timing (DNA abundance) of each origin in the mutant vs. wild type. The group of origins predefined in previous experiments as regulated in the respective mutant is color-coded. The distributions of residual changes, following normalization by the predicted replicon length, are also shown (lower panel). P-values are measured based on a two-sample t-test (or one-way ANOVA in the case of the number of groups > 2) comparing the distribution of residuals in the regulated vs. unregulated origin groups. Origin classification is based on the following references: fkh1,2ΔΔC and fkh1Δ taken from Knott et al. (2012); rpd3Δ and sin3Δ were classified based on the up-regulated origins in rpd3Δ from Knott et al. (2009); ctf8Δ and ctf18Δ classification was taken from Crabbe et al. (2010). Classification of clb5Δ origins was taken from McCune et al. (2008). Subtelomeric origins in yku80Δ, rif1Δ, sir2Δ, sir3Δ, and sir4Δ were defined within 50 kb from the telomeres (Methods). (C) Replication profile around centromeres. Shown are the replication profiles around the centromere (marked in a dashed red line) of Chromosome 16 for the indicated strains (green) compared to the wild-type strain (black). (D) The unexplained profile score correlates with local changes. For the indicated strains, the unexplained percentage of the signal (x-axis) is compared to the significance of change in centromere replication. Significance is defined by the −log10(P-value) of a t-test comparing changes in replication time of centromeric vs. noncentromeric origins (y-axis). (E) DNA abundance around centromeric regions. Shown are 10-kb windows around the centromere for the indicated strains.











