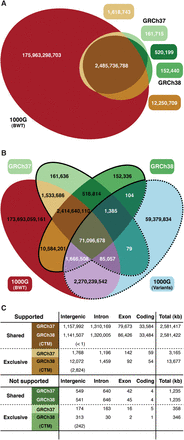
(A) 31-mer intersection of two human reference assemblies (GRCh37 and GRCh38) and the 1000GP population BWT. (B) 31-mer intersection of two human reference assemblies, 1000GP population BWT, and all 31-mers generated from the 1000GP phase 3 SNP and indel variants (The 1000 Genomes Project Consortium 2015). 31-mers shared between reference sets and variant set (white numbers) make up for ∼3% of each data set and almost all (99.998%) are supported by the 1000GP population BWT. (C) A breakdown of the regions on the two human assemblies with and without 1000GP population BWT support that are shared or exclusive to either genome build (all numbers are kbp), in four functional categories. (CTM) Centromeric sequence.











