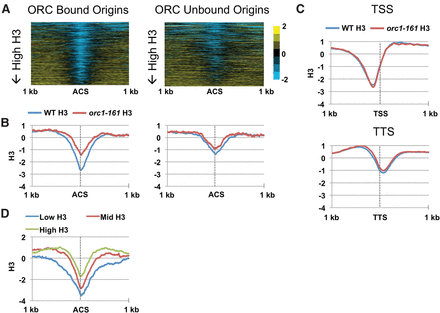
Nucleosome occupancy varies across origins in G1 and is ORC-dependent. (A) Nucleosome occupancy (log2) in G1 aligned around the ACS for 798 predicted yeast origins from the OriDB database (Nieduszynski et al. 2007). Origins are classified based on ORC binding (397 ORC bound and 401 ORC unbound). Origins were aligned from high (top) to low (bottom) nucleosome occupancies. (B) Average nucleosome occupancy after Z-score normalization across all ORC-bound (left) and ORC-unbound origins (right) spanning 1 kb upstream of and downstream from the ACS. The dotted line represents the midpoint of the ACS. WT nucleosome occupancy is shown in blue, and orc1-161 (at a permissive temperature) H3 is in red. (C) Average nucleosome occupancy 1 kb upstream of and downstream from 4551 transcription start sites (TSSs) and 5120 transcription termination sites (TTSs) (Nagalakshmi et al. 2008). (D) Average nucleosome occupancy of ORC-bound origins (393) classified based on average H3 levels 500 bp upstream of and downstream from the ACS: Low nucleosome occupancy includes the bottom 33% (blue), mid H3 includes the middle 33% (red), and high H3 includes the top 33% (green).











