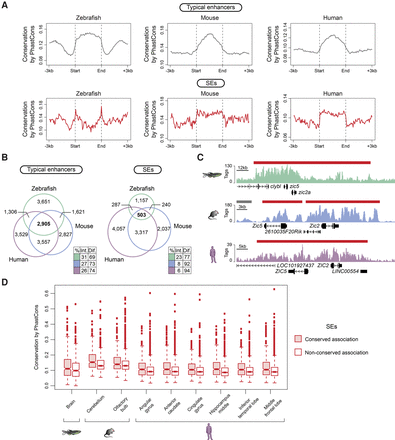
SE conservation in vertebrates. (A) Metagenes of sequence conservation of typical enhancers and SEs from zebrafish whole brain, mouse olfactory bulb, and human middle frontal lobe. The x-axis depicts the start and end of typical enhancers and SEs flanked by 3 kb of adjacent sequence. The y-axis represents sequence conservation calculated by PhastCons. (B) Venn diagrams show the number of orthologous genes associated with brain typical enhancers (left) and SEs (right) in zebrafish (green), mouse (blue), and human (purple). Color-coded tables show the percentages of intersection and difference for each species. The observed differences in overlap between typical enhancers and SEs in the three species are significant (P-values ≤5.497 × 10−8) based on G-tests of independence. (C) ChIP-seq binding profiles for H3K27ac at the indicated loci in zebrafish, mouse, and human brain (raw tag counts represented on the y-axis). Typical enhancers and SEs are denoted by gray bars and red bars, respectively. Gene positions are noted along the x-axis. (D) Box plots depicting average sequence conservation of brain SEs with maintained orthologous association in zebrafish, mouse, and human and with no maintained orthologous association. The y-axis shows sequence conservation calculated by PhastCons. The box bounds the interquartile range divided by the median, and the notch approximates a 95% confidence interval for the median. All observed differences in conservation between SE categories are significant (P-value ≤9.1 × 10−3) based on Wilcoxon rank-sum tests.











