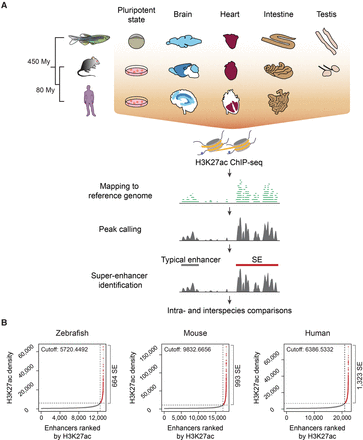
Identification of typical enhancers and SEs in vertebrate genomes. (A) Workflow for the identification of vertebrate typical enhancers and SEs. Schematic representations depict the cells and tissues analyzed. (B) Saturation curves of H3K27ac density across brain data sets (whole brain for zebrafish, olfactory bulb for mouse, and middle frontal lobe for human). The number of ranked typical enhancers and SEs by H3K27ac density (x-axis) and their densities (y-axis) are plotted. Horizontal dotted lines represent density cutoffs used for the classification of SEs and vertical dotted lines demark SEs from typical enhancers. The total number of predicted SEs is noted on the right side of each graph.











