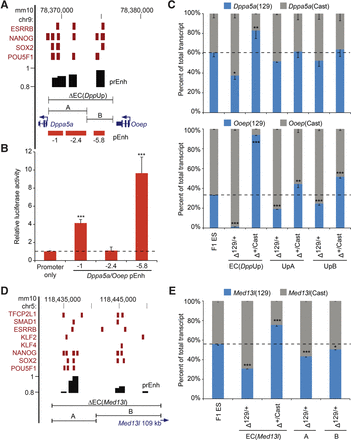
Partial enhancer cluster deletions reveal reduced effects on gene expression compared to larger deletions. (A,D) Schematic representation of Dppa5a/Ooep (A) and Med13l (D) EC regions. Transcription factor-bound regions (red bars) obtained from the CODEX database are shown. Predicted enhancers (prEnh) are shown in black. The deleted enhancer cluster (ΔEC) and partial deletions (A,B) are shown with a line that links the 5′ and 3′ gRNA targets. All data are displayed on the mm10 assembly of the UCSC Genome Browser. (B) Enhancer activity was identified in a reporter assay for the −1 enhancer and the −5.8 enhancer within the EC(DppUpA) and EC(DppUpB) regions, respectively. Error bars represent SEM. A significant difference from the promoter only vector is indicated: (***) P < 0.001. (C,E) Partial deletion of an EC has a reduced effect on expression of the linked Dppa5a/Ooep (C) or Med13l (E) allele compared to the full deletion. Allele-specific primers detect 129 or Cast RNA in RT-qPCR from F1 ES, Δ129/+, and Δ+/Cast clones. Expression for each allele is shown relative to the total. Error bars represent SEM. Significant differences from the F1 ES values are indicated: (*) P < 0.05, (**) P < 0.01, (***) P < 0.001.











