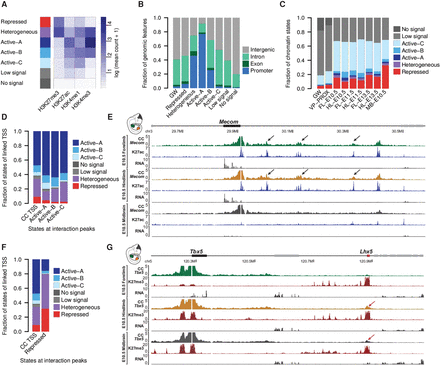
Interactions associate with functional chromatin states. (A) EpiCSeg segmentation from H3K4me3, H3K4me1, H3K27ac, and H3K27me3 generates three types of active states, one repressed state, a heterogeneous state, and two types of states depleted from the surveyed chromatin marks. (B) Distribution of genomic features across chromatin states in hindlimb at E10.5, as well as their genome-wide distribution. (C) Functional chromatin domains cover 19% of the genome-wide (GW) and 25% of regions proximal to the CC viewpoints (±2 Mb) (VP-PROX). More than sixty percent of interaction sites (99th percentile) were found in functional chromatin segments. (D) States of viewpoints’ TSSs linked to active interaction peaks (data were aggregated from all tissues) (see Supplemental Fig. S7A). (E) At the Mecom gene, interaction peaks form specifically over acetylated regions (black arrows) and disappear in the midbrain, where these regions are not acetylated. The RNA-seq tracks show that the Mecom gene is expressed at high level in limb tissues, whereas its expression is much lower in midbrain. (F) States of viewpoints’ TSSs linked to repressed interaction peaks (data were aggregated from all tissues) (see Supplemental Fig. S7B). (G) Tbx5 promoter forms a long-range interaction (red arrow) with an H3K27me3 domain (Lhx5) only when it is covered by the same mark and repressed (see RNA tracks) in hindlimb and midbrain.











