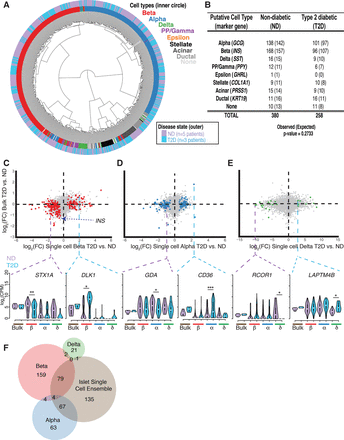
Single-cell transcriptome analyses identify cell-type–specific expression changes in T2D islets. (A) T2D and ND single-cell transcriptomes cluster together by cell type after unsupervised hierarchical clustering. (B) Number of each ND and T2D cell type classified by marker gene expression as shown in Figure 2. The numbers of cells expected in each condition based on a χ2 test are indicated in parentheses. (C–E, top) Scatter plots of log2 fold-change (FC) expression detected between T2D and ND samples from bulk intact RNA-seq (y-axis) and from Fluidigm C1 single-cell RNA-seq (x-axis) from beta cells (left plot; red), alpha cells (middle plot; blue), and delta cells (right plot; green). (Bottom) Violin plots highlight examples of differentially expressed genes in one single-cell type. Dashed purple lines represent repressed genes in the respective T2D cell type, while dashed blue lines represent induced genes. (*) FDR < 0.05, (**) FDR < 0.01, (***) FDR < 0.001. (F) Venn diagram showing the intersections of differentially expressed genes identified between T2D and ND transcriptomes at single-cell-type and islet single-cell ensemble resolution. The islet single-cell ensemble represents the pooled collection of beta, alpha, delta, and PP/gamma single cells.











