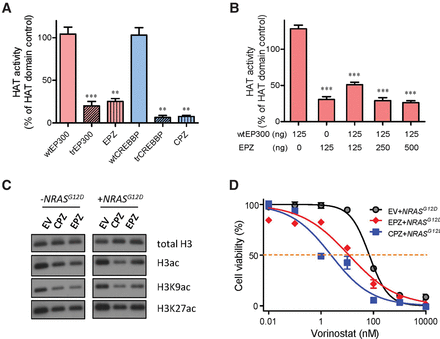
EP300- and CREBBP-ZNF384 fusions resulted in loss of HAT activity, global histone acetylation deregulation, and sensitivity to HDAC inhibition. (A,B) HAT enzymatic activity was measured for various EP300 and CREBBP proteins: wild-type, truncated, and ZNF384 fusions. Proteins were expressed in Sf9 insect cells and purified to homogeneity. (A) HAT activity was determined for each protein individually. (B) In parallel, wild-type EP300 was mixed with increasing amount of EP300-ZNF384 fusion protein to examine dominant-negative effects of the latter on HAT activity. (C) Global H3, H3K9, and H3K27 acetylation were evaluated by Western blot in Ba/f3 cells overexpressing EP300-/CREBBP-ZNF384 fusion, with total H3 as a loading control. (D) Cytotoxicity of HDAC inhibitor vorinostat was examined in NRASG12D-transformed Ba/f3 cells expressing EP300-ZNF384 (red) or CREBBP-ZNF384 (blue) or transduced with empty vector (gray). Cells were incubated with vorinostat for 48 h, and viability was then measured using a MTT assay. Experiments were performed in triplicate and repeated at least three times. (wtEP300) Wild-type EP300; (trEP300) truncated EP300; (wtCREBBP) wild-type CREBBP; (trCREBBP) truncated CREBBP; (EPZ) EP300-ZNF384 fusion; (CPZ) CREBBP-ZNF384 fusion; (EV) empty vector. EPZ- or CPZ-transduced cells were significantly more sensitive to vorinostat than EV, as determined by two-way ANOVA (P < 0.001). (A,B) Statistical significance was evaluated by using a two-sided Student's t-test: (**) P < 0.01; (***) P < 0.001.











