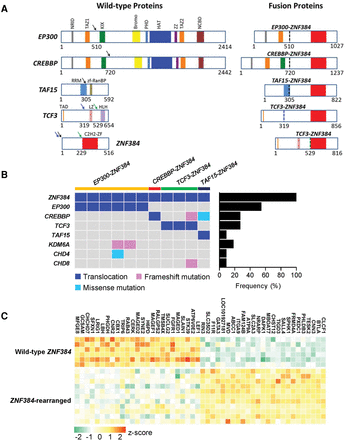
Genomic abnormalities in ZNF384-rearranged ALL. (A) Schematic representation of ZNF384 rearrangement in ALL. (Left) The wild-type proteins and their domain structures. Arrows indicate breakpoints observed in ALL samples, and colored blocks represent protein functional domains. (Right) The predicted ZNF384 fusion proteins. Dashed vertical lines mark the junction between fusion partners and are color-coded according to the breakpoint (arrows) in the left panel. (B) ZNF384-rearranged cases were also subjected to whole-exome sequencing, and putative functional genomic lesions (translocation, frameshift, and missense mutations) in selected genes are indicated for each case (left). (Right) Bar graph denotes the cumulative frequency of genomic lesion in each gene in ZNF384-rearranged ALL. (C) Heatmap shows the top 50 differentially expressed genes. Columns indicate genes, and rows are patients or patient groups. Expression value is indicated for individual patient with ZNF384 rearrangement; for cases without ZNF384 rearrangement (wild-type), expression is shown as the mean expression value for each ALL subgroup. The normalized expression level for each gene (z-score) is indicated by a color (red and green for over- and underexpression, respectively).











