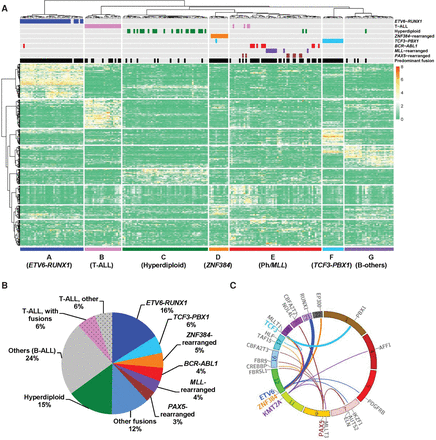
Whole-transcriptome sequencing of childhood ALL. (A) Unsupervised hierarchical clustering of global gene expression profile from 231 children with newly diagnosed ALL. Columns indicate ALL patients, and rows are genes; gene overexpression and underexpression are shown in red and green, respectively. ALL immunophenotype, cytogenetics, and selected chromosomal translocations are indicated for each sample above the heatmap, and the text below denotes seven major ALL subgroups identified on the basis on gene expression profiles. “Predominant fusion” refers to likely functional and driver fusion event, as defined in the Methods and Supplemental Figure S3A. (B) Pie chart of major gene fusions and cytogenetic abnormalities identified in B-ALL and T-ALL. Percentage indicates prevalence in the entire cohort; B-ALL and T-ALL are discriminated by solid fill and dot pattern, respectively. (C) Circos plot of the predominant fusions involving frequently translocated genes (i.e., genes with three or more different translocation partners identified). The line links the two partner genes in a fusion, with line width indicative of the prevalence of the specific fusion.











