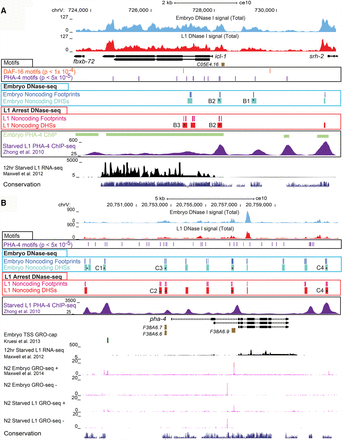
L1 arrest noncoding DHSs discovered in genes up-regulated during L1 arrest. Total DNase signal (red) from both strands of L1 arrest DNase-seq and embryo DNase-seq (light blue) are shown. L1 arrest noncoding DHSs (red boxes) and associated TF footprints (pink boxes), as well as embryo noncoding DHSs (light blue boxes) and associated TF footprints (dark blue boxes), were detected. Asterisks indicate DHSs that were tested for activity in transgenic C. elegans (Supplemental Table S9). Additional tracks shown are C. elegans RefSeq genes (black boxes with arrows), noncoding transcripts (brown boxes), and 12 h starved L1 mRNA-seq tracks (black) from Maxwell et al. (2012), phyloP sequence conservation (dark blue) are also shown. Other comparison tracks include PHA-4 ChIP-seq from embryo (light green) and starved L1 larvae (purple) (Zhong et al. 2010). PHA-4 (purple boxes) and DAF-16 (orange boxes) motifs, as well as TSS (L1 starved GRO-cap data from Kruesi et al. 2013 as dark green boxes; and L1 and embryo GRO-seq data from Maxwell et al. 2014 as magenta signal), are shown when relevant. (A) Noncoding DHSs of icl-1. One L1 arrest–associated noncoding DHS containing a DAF-16 binding motif (P < 1 × 10−4 threshold) and a PHA-4 motif (P < 5 × 10−5) is detected in the first intron of icl-1. TFs footprints are found within this DHS that overlap the DAF-16 motif. Three other noncoding DHSs are detected in both L1 and embryo coinciding with PHA-4 ChIP-seq peaks from L1 starved larvae (Zhong et al. 2010), and two of them harbor TF motifs. Two additional upstream regions bound by PHA-4 in L1 starved larvae were not detected. (B) Known and novel CRMs of pha-4. Four embryo and three L1 arrest noncoding DHSs are observed upstream of the longest transcript, pha-4a. One of these is an embryo-associated noncoding DHS overlapping an TSS that was detected in embryos but not L1 arrest by GRO-cap (data from Kruesi et al. 2013). Directly upstream of pha-4a is an L1 arrest–associated noncoding DHS that overlaps PHA-4 TF binding sites. The two noncoding DHSs upstream of C1 were tested in one transgenic construct, but unlike other DHSs tested in the locus, it did not drive expression.











